Retrieval Transformers for Medicine
Exploring how DeepMind's RETRO can benefit health and medicine
Deep learning has a weight problem. Large language models such as GPT-3, Megatron-Turing, and Gopher are a significant step towards foundational models capable of performing language tasks at human level. Although impressive in their abilities, these models are unwieldy.
The 175 billion parameter GPT-3 requires 20-25 GPUs for inference alone1. If the inference cost is not a deterrent, the compute cost to train large language models can easily run in the 10s of millions of dollars. The issue of cost in training and inference for these models has resulted in few groups applying large language models to biomedicine, despite the huge promise they have shown in general NLP tasks.
One optimistic school of thought is that the issue of compute and cost will be sorted out by Moore’s law2. After all, the relentless decrease in GPU compute cost over the last 10 years is what made these large language models possible in the first place. However, until the day when Moore’s law has caught up with the size of these foundational models, there is considerable pressure to achieve the same performance level with much smaller models.
Just recently, it appears that DeepMind have demonstrated one such contender, named RETRO or Retrieval-Enhanced Transformer.
In this post I will briefly explain what makes RETRO special, and how these properties are particularly suited to machine learning in health and medicine.
What’s so special about RETRO?
To understand RETRO, I highly recommend Jay Alammar’s The Illustrated Retrieval Transformer. In fact, while you are visiting his blog, read everything. In short, the main difference between previous large language models and RETRO is in where the majority of knowledge3 is stored.
Both “vanilla” transformers such as GPT-3 and retrieval-enhanced transformers (RETRO) are trained on huge language datasets. For GPT-3, all knowledge about the dataset must be stored in the neural network’s weights. The working theory for large language models is that larger models have a greater capacity to store this knowledge, as well as a greater capacity to process the relationship between the pieces of knowledge. Part of what makes GPT-3 effective is that the model is so large that a significant portion of the available knowledge within the training dataset can be captured within the model weights. In contrast, RETRO attempts to separate the knowledge from higher-order processing of that knowledge by storing the knowledge in a separate database.
RETRO has two main components. A transformer-based neural network and a database.
The database is a key-value store. The key is an embedding vector and the value is a corresponding piece of text. The key or embedding vector is generated by applying a neural network (for RETRO, they used BERT) to the text to generate a vector representation. For a given training dataset, all text snippets from the dataset are encoded and stored in the database. As a result, the database contains all the knowledge from the training dataset.
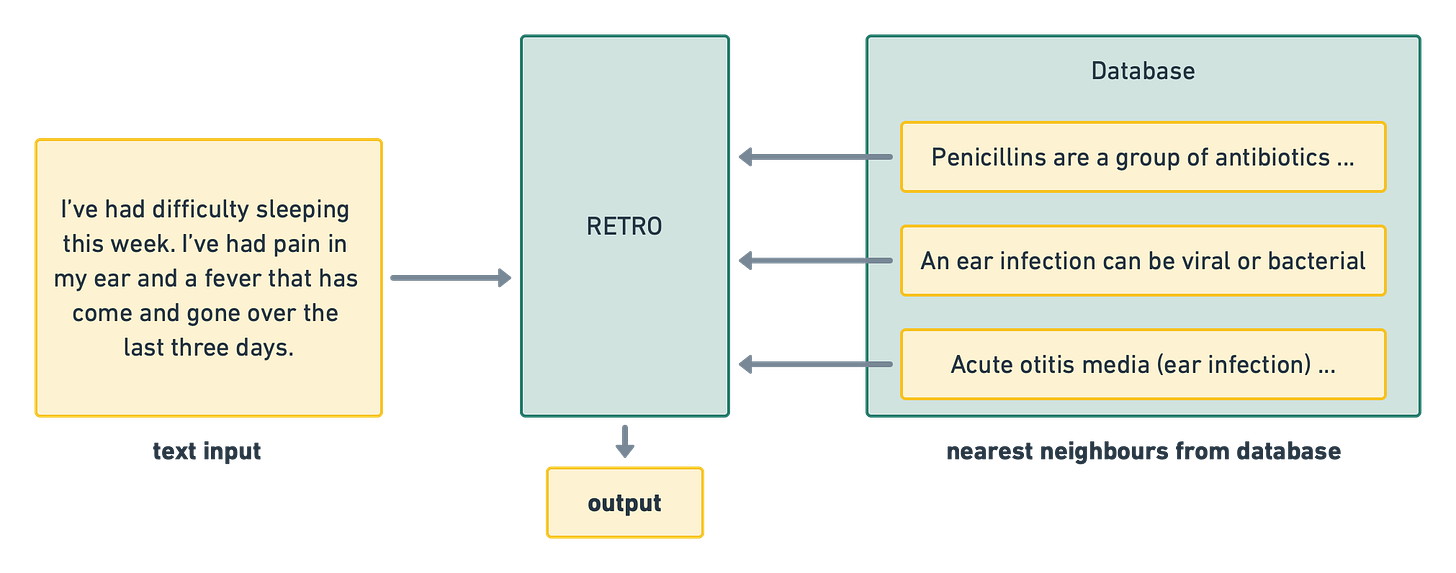
The transformer part of RETRO combines the input with knowledge from the database. When an input is fed into RETRO, this input is encoded using BERT. Next, the encoded input is used in a nearest-neighbour search in the database. This step allows RETRO to find the most closely related pieces of knowledge stored in the database. The result of the nearest-neighbour search is then fed into the transformer network alongside the original input. As a result, the transformer portion of RETRO is able to incorporate important snippets of related knowledge without having to memorize this information within the neural network weights.
Disentangling knowledge about the world from other aspects such as language knowledge opens up the possibility to scale world knowledge independently. An added benefit is that RETRO’s database can be expanded without retraining, since the neural network used to encode the database text is fixed and separate from the RETRO neural network.
There are many other nuances and improvements from the RETRO paper, but the core idea is just that!
RETRO in medicine
RETRO will likely impact multiple areas of machine learning including NLP, computer vision and audio processing. Here I’ll focus on NLP in healthcare, using a toy example to show the advantages built into RETRO.
Imagine you are building a chatbot to function as a conversational WebMD. Users will come to your chatbot with a description of their symptoms and medical history. The task for the chatbot is to make a diagnosis and recommend a treatment.
Patient: I’ve had difficulty sleeping this week. I’ve had pain in my ear and a fever that has come and gone over the last three days.
Info: Male, 29 years old, no history of medical issues, no allergies.Dr Chatbot: You have an ear infection, we recommend that you are prescribed penicillin, an antibiotic that will help you fight the infection.
We can break down the task into the following components:
Extract symptoms
Extract medical history
Match symptoms and medical history to a diagnosis
Match diagnosis to a treatment
Generate text reply describing the diagnosis and treatment
We are able to train this bot in an end-to-end manner. We have to collect a dataset that consists of patient-doctor chat transcripts. This could be curated data from existing Telehealth chat services or a dataset generated specifically to train our model. Outside of the biomedical field, there are many chatbots (“conversation engines”) trained in such a way and deployed in production at this very moment.
RETRO and new knowledge
Now let’s imagine that we deploy our biomedical chatbot. Six months after deployment, a new treatment, Newcillin, is developed for ear infections. Newcillin is highly effective, but only to be used in children aged 5 to 12 years old. We now need to update our chatbot and underlying machine learning model with this new treatment. For traditional transformers, this would require updating the training dataset. We would have to take care to ensure there are sufficient training samples that mention Newcillin, and then fine-tune the model or completely retrain end-to-end.
For a large language model such as GPT-3, this would be a costly and difficult process. Retraining every time a new treatment or discovery is made is impractical. Fine-tuning, although more practical, can lead to catastrophic forgetting and loss of performance over time.
For a retrieval-enhanced transformer, the process to update the model with new knowledge can be as simple as adding the relevant knowledge to the database. If we are able to generate text snippets that represent the new information, we can generate their corresponding embeddings and add these key-value pairs to the database.
RETRO and attribution
In medicine, we often want to explain decisions in the context of evidence. RETRO’s database and nearest neighbour search provides a built-in glimpse at the snippets of knowledge that contribute towards the output. This results in a more explainable machine learning model, useful for interpreting the output and understanding cases where the output is wrong.
Compare this to other transformers, where it is often not possible to attribute the output to specific snippets of text from the training dataset. When training a large language model, we assume that the majority of knowledge from the training dataset is captured within the model. However, since knowledge and higher order processing are entangled, we cannot easily determine which pieces of knowledge are accurately captured.
RETRO and amending knowledge

Like all models, retrieval model performance is affected by the quality of the training dataset. If there are many errors and factual inaccuracies in the training dataset, the resulting model will be worse. Unlike other models, retrieval models provide a simple way to amend the knowledge at inference time. If an incorrect output is observed, we can take a look at the retrieved text, amend any false statements, update the database and then re-run inference.
Conclusion
NLP has had a huge impact across the tech industry over the last 10 years. Transformers power modern search engines, recommendation systems, chatbots and much more. Comparatively, NLP’s impact within health and medicine has been much smaller. Partially, this has been caused by lack of interpretability and unusual failure modes in existing models, limiting their suitability for medicine. Although I’m just scratching the surface here on retrieval models, I hope this post highlights the promise they bring for powering the next generation of health software.
GPT-3 was trained with bfloat16 precision.
1.75B parameters = 350GB of VRAM required
Assume 16GB GPU VRAM
= minimum of 21.875 GPUs (assuming zero overhead)
When discussing GPUs, some people refer to this as Huang’s law
I’m using the word knowledge carelessly here. All state of the art NLP models are just algorithms that predict the most probable word (as in highest probability), given an input, such as the previous words in a sentence. In many cases when interacting with and using large language models, the output can lead you to believe that the generating algorithm has knowledge and perhaps even understanding about the topic at hand. We’re teetering on the edge of philosophy at this point. It’s not my place to say whether a neural network can possess understanding and knowledge about language. I can confidently say that a neural network can contain information that maps very closely to knowledge. In this context, that is what I mean by knowledge.